Advancing Bioinformatics Through Computational Innovation
My research develops machine learning methods, bioinformatics tools, and data resources to analyze complex biological systems. I focus on genomics, transcriptomics, and host-pathogen interaction modeling, with an emphasis on building practical tools that are interpretable and reproducible.
I collaborate across disciplines to translate algorithms into accessible web resources and pipelines. My current work at South Dakota State University centers on AI-enabled discovery in agriculture and animal health, integrating multi-omics data to improve prediction and biological understanding.
Core themes and scientific focus
Four pillars guiding ongoing investigations, tool development, and collaborative projects.
Multi-omics Data Integration
Developing computational methods to merge genomics, transcriptomics, proteomics, and metabolomics for a systems-level view of biology.
- • Algorithms for multi-omics data fusion
- • Machine learning for cross-platform analysis
- • Visualization of high-dimensional biological data
- • Statistical modeling of integrated signals
AI in Genomics
Applying deep learning to predict gene function, regulatory elements, and variant effects with interpretable, robust models.
- • Deep learning models for genomic prediction
- • Interpretable AI for sequence analysis
- • Variant prioritization and functional scoring
- • Precision medicine applications
Systems Biology
Modeling biological networks and pathways to uncover mechanism-level insights and identify intervention points.
- • Network analysis of biological systems
- • Metabolic pathway modeling
- • Gene regulatory network inference
- • Systems-level target discovery
Bioinformatics Tooling
Designing databases, web servers, and analysis pipelines that make complex data usable for researchers.
- • NGS workflows and automation
- • Specialized biological databases
- • Scalable backends and APIs
- • User-focused interface design
Highlighted research platforms
Selected tools and resources spanning deep learning, RNA analysis, and host-pathogen interactions.
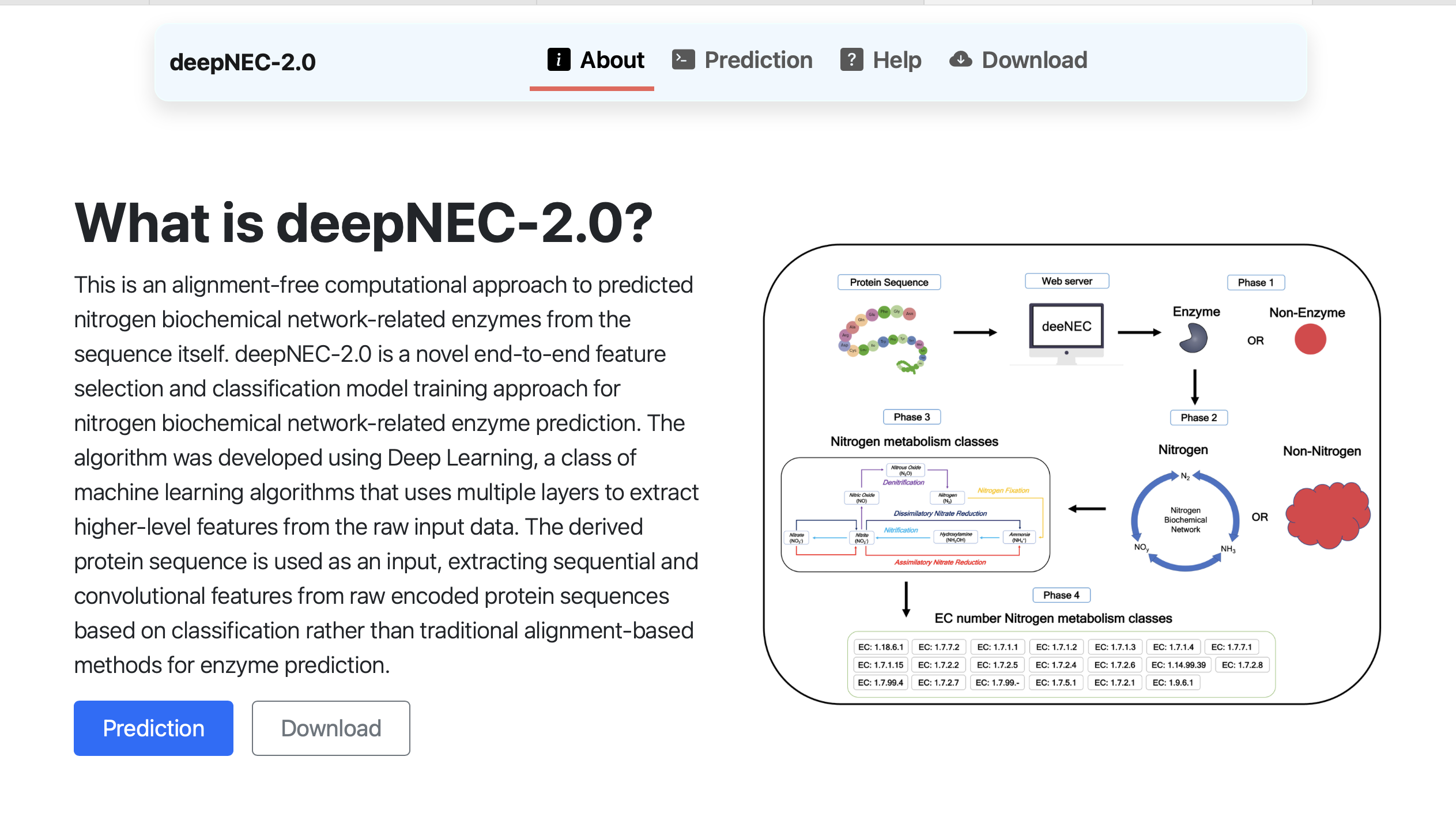

deepNEC
A deep learning framework for predicting nitrogen metabolism enzymes from protein sequences, providing interpretable insights into sequence-function relationships.
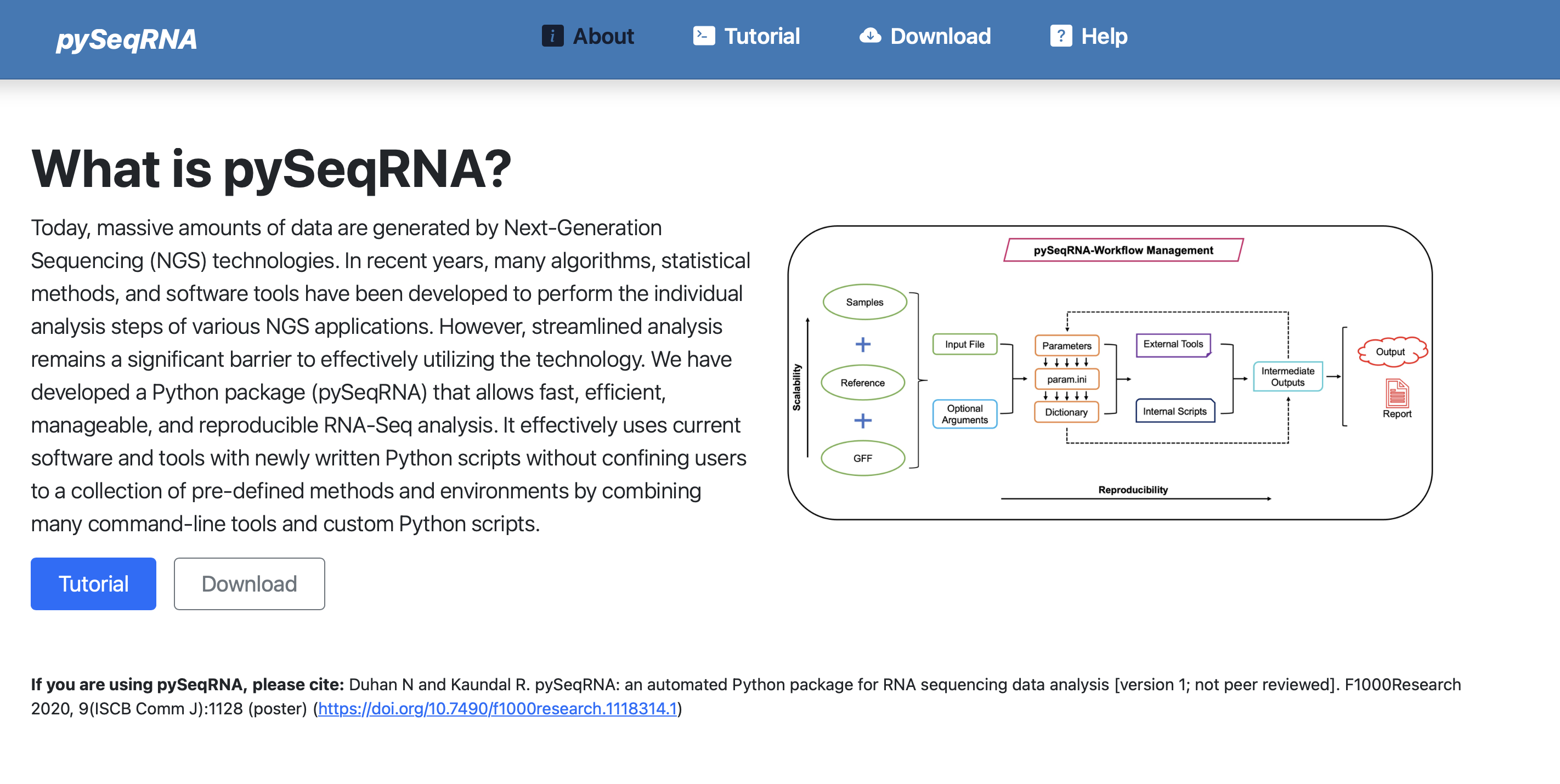
pySeqRNA
A comprehensive RNA-Seq analysis package with automated QC, alignment, quantification, and differential expression workflows.
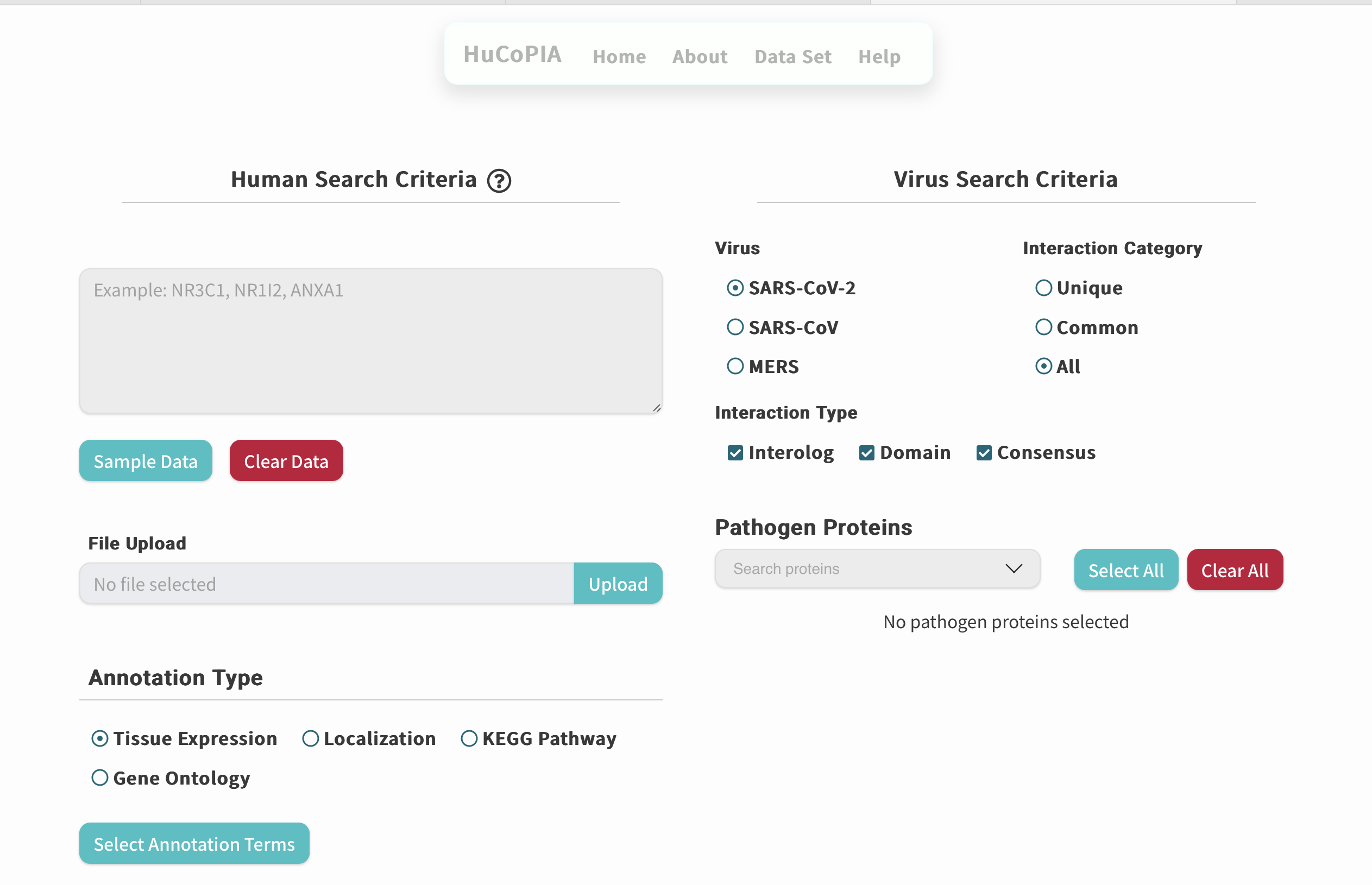
HuCoPIA
An atlas of human-coronavirus protein interactions with network visualization and comparative analyses across Coronaviridae.
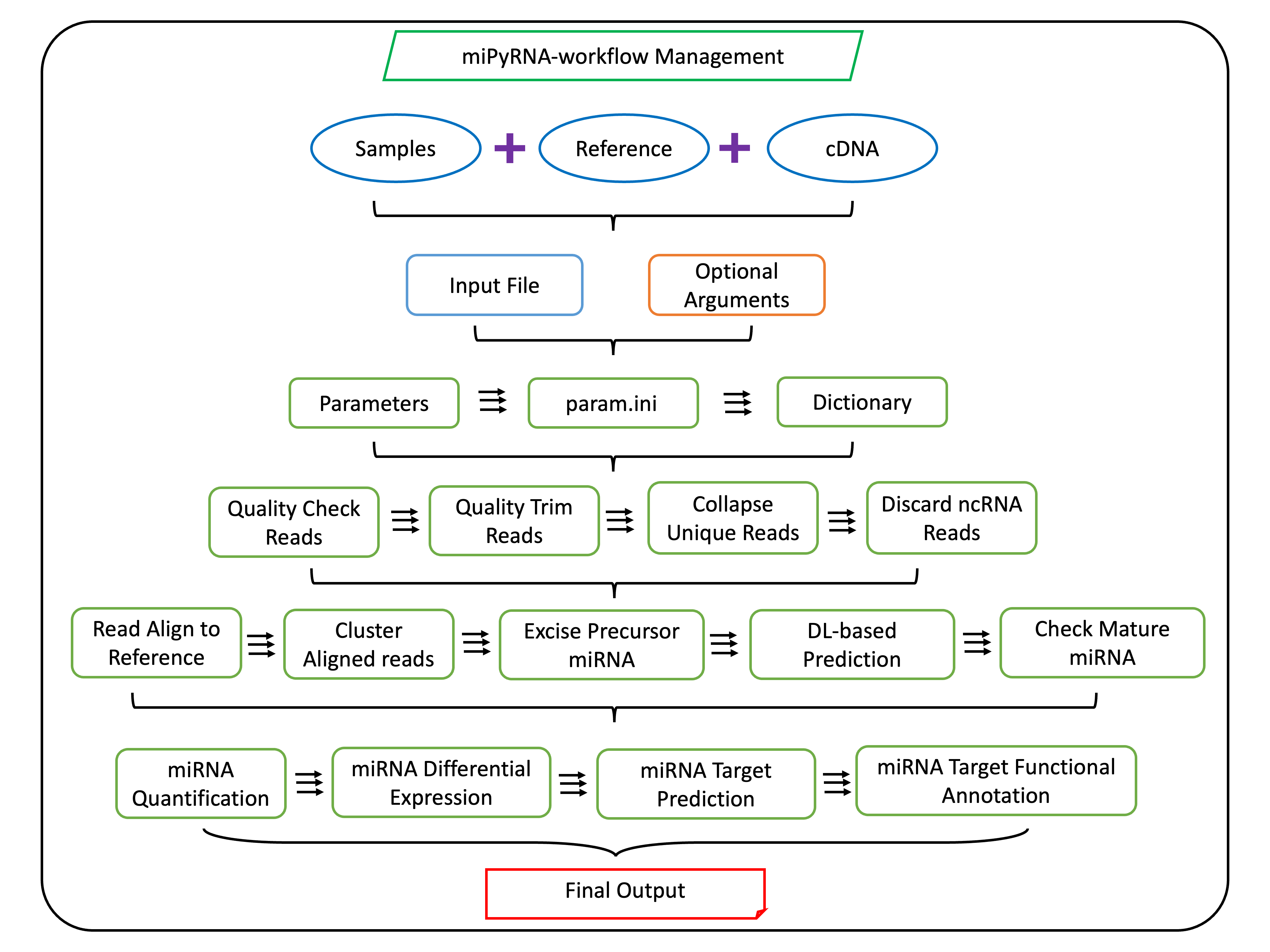
miPyRNA
A Python package for small RNA-seq analysis focused on microRNA identification, quantification, and target prediction.
Open to collaborative research and tool development
If you are working on genomics, host-pathogen interaction modeling, or AI-driven bioinformatics, I would love to collaborate.